
Dualase genome editors have two sites that leave different DNA ends to bias natural cell repair pathways to the correct genome editing outcomes
®

Dualase editors are easily programmed without compromising high editing efficiency and precision with their differentiated features:
®
(1) The I-TevI domain only cuts target DNA if bound to correct Cas site and properly spaced from Cas site
(2) Dualase has high specificity through a proprietary engineered Cas domain and its two-site requirement
(3) Programmable through targeting RNA & the I-TevI target site 'CNNNG' occurs frequently in the genome for high programmability
(4) Dualase leaves different DNA ends at the I-TevI and Cas sites: a 'sticky' end and a 'blunt' end
(5) No detectable off-target edits in vitro or in vivo using targeted and genome-wide surveys - proprietary engineered Cas domain and two-site requirement
®
®
Correction of insertions, deletions and base change mutations with a small, highly deliverable cargo for in vivo editing





®

Versatile and programmable Dualase genome editors are small (~3.7kb) and fit into a single AAV or are deliverable as small RNAs in lipid nanoparticles and polymers
Dualase genome editors use distinct mechanisms of action for sequence repair or removal
®
Mechanism 1 Dual-guided I-TevI domains for precise sequence removal
Dual-guided I-Tevl for efficient & precise large sequence removal, including large repetitive sequences, and deliverable as single adeno-associated viral vectors (AAV) or small RNAs

Large sequence
Inactive Cas

(1) Dual guides bind outside of a large sequence insertion and orient a version of Dualase with an inactive Cas domain and an active I-TevI domain to either side of a large sequence
®
(2) Cutting at both I-TevI domains removes the large sequence and leaves two complementary 'sticky' ends
(3) The two complementary ends are joined through a preferred non-homologous end-joining pathway (NHEJ)
(4) Large (kilobases) sequences are precisely removed
Advantages of Dualase precise sequence removal
Deliverability
Dualase and dual guides can be packaged into a single adeno-associated viral vector (AAV) to deliver to many tissues, including CNS
Efficiency
Sticky 3' DNA ends are efficiently repaired using the ubiquitous non-homologous end-joining (NHEJ) pathway
Precision
Scarless repair ensures no nearby important sequences are modified
Versatility
Capable of repairing large sequence mutations, including repeat expansions, to leave non-pathogenic numbers of repeats
®
®
Mechanism 2 Repair or insertion of small or large sequences
Dualase uniquely uses a common, naturally-occurring repair pathway for RNA-templated DNA repair
®
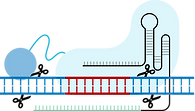
(1) I-TevI & Cas cutting



Pol θ
(2) Sequence between I-TevI & Cas site removed
(3) Free end of RNA repair template binds to 'sticky' end of I-TevI site
(4) Naturally occurring polymerase theta (θ) recruited to copy RNA sequence into the genome
RNA-templated DNA repair for small (100's of bases) and large (1000 bases) sequences
Advantages of Dualase repair
Deliverability
All components encoded in as a single DNA or RNA molecule (no complex payloads)
Small Size
No need for other large repair proteins (e.g., reverse transcriptase, base editor or recombinase)
Precision
High fidelity scarless repair
Versatility
Repairs all mutation types (insertions, deletions and single base changes)
Efficiency
Capable of precise kilobase sequence insertions